The era of genomic medicine has reached a tipping point. Thousands of healthcare professionals in the US and around the world are using the concepts and principles of the “omics” revolution to personalize prevention and treatment strategies. By Joe Veltmann PhD FAAIM DCCN and Roberta (Bobbi) Kline MD FACOG
Every month, it seems, the suffix “omics” is added to another established branch or field of scientific study: pharmacogenomics, nutrigenomics, metabolomics, toxicogenomics, lipidomics and microbiomics.
While you may not be using genomic testing in your clinical practice today, more and more patients want their healthcare professional to interpret direct-to-consumer (DTC) genomic test results, to validate their internet searches and to individualize pharmacological prescriptions, dietary and nutritional recommendations, exercise regimens and prevention programs to ward off degenerative diseases associated with aging.

Unlike many of the medical innovations of the past where medical devices, procedures and information were phased in over time, allowing healthcare professionals sufficient time to assimilate the new information and determine how best to use it in his/her clinic, genomic medicine has been traveling at warp speed ever since the completion of the Human Genome Project (HGP) in 2003.
Today, healthcare professionals are confronted with massive amounts of genomic information from DTC firms, new terminology and, at the same time, trying to reconcile the genetic information they learned in medical or graduate school with this new scientific discipline.
Our informal conversations with healthcare professions across many disciplines confirm what survey after survey has revealed. The majority of healthcare professionals do not feel comfortable discussing genomic test results with their patients, feel unprepared to advise them, and are confused about how to interpret the results. As a result, clinicians oftentimes dismiss all genomic test results as “junk”, regardless of the source.
The intent of this article in Today’s Practitioner and those to follow is to clear up any confusion healthcare professionals might have about the differences between genetics versus genomics, nutrigenetics and nutrigenomics and how genomic testing can unravel nutritional paradoxes to personalize and more accurately determine a patient’s nutrient requirement.
Today, let’s clarify the difference between genetics and genomics.
Although genetics and genomics sound alike, and are often used interchangeably in articles and webinars, important scientific and clinical distinctions exist between these two scientific fields of study.
“Genomics is the ‘new kid on the block’ compared to genetics.” The classical definition of genetics is the study of heredity – or how the characteristics and traits (phenotypes) of a living organism are transmitted from one generation to the next. This occurs via deoxyribonucleic acid (DNA), a double helix molecule in the cell’s nucleus that comprises genes–the basic unit of heredity. You may recall that many of the early principles and rules of heredity were discovered by an Augustinian monk and scientist, Gregor Mendel. His seminal research with pea plants in the mid-1800s laid the foundation for modern-day genetics.
Genomics is the “new kid on the block” compared to genetics, and became possible only in the last couple of decades due to technical advances in DNA sequencing and computational biology. In 1976, Belgian scientists succeeded in fully sequencing the genome of bacteriophage MS2, a bacteria-infecting virus. They identified all 3,569 DNA base pairs — the 4 nucleotides (Adenosine, Cytosine, Guanine and Tyrosine) — that make up the DNA code. The first animal genome was completed in 1998. Five years later, more than 1,000 researchers from six countries collaborated on the HGP and identified all 3.2 billion DNA base pairs in the human genome.
One of the biggest takeaways from the HGP was that humans share 99.5% of their genome with each other. However, the small differences are quite significant–they account for all the different characteristics and traits (phenotypes) that exist among humans: hair or eye color, height, weight, skin pigment, disease risk, etc.
When genomics is defined in the context of genetics, it is the study of the entirety of an organism’s genes – called the genome.
Genomic researchers analyze enormous amounts of DNA-sequence data to find variations that affect health, disease or drug response. In humans, that means searching through about 3.2 billion units of DNA across 23,000 genes. Translating the research data into usable clinical information can be daunting and represents an ongoing task.
In a clinical sense, genetics is the study of single genes or parts of genes and their effects on a person’s 
Rather than segregating them into discreet entities, more and more scientists see genetics and genomics as part of a continuum. On one end of the spectrum, there are rare, single gene disorders in the germ line with high penetrance. On other end, there are common, low penetrance gene variants or single nucleotide polymorphisms from multiple locations interacting with environmental factors, leading to complex diseases.
Don’t Miss East Meets West Podcast: Genomic Testing and Personalized Health: Getting Started with the Basics, Click Here to Listen
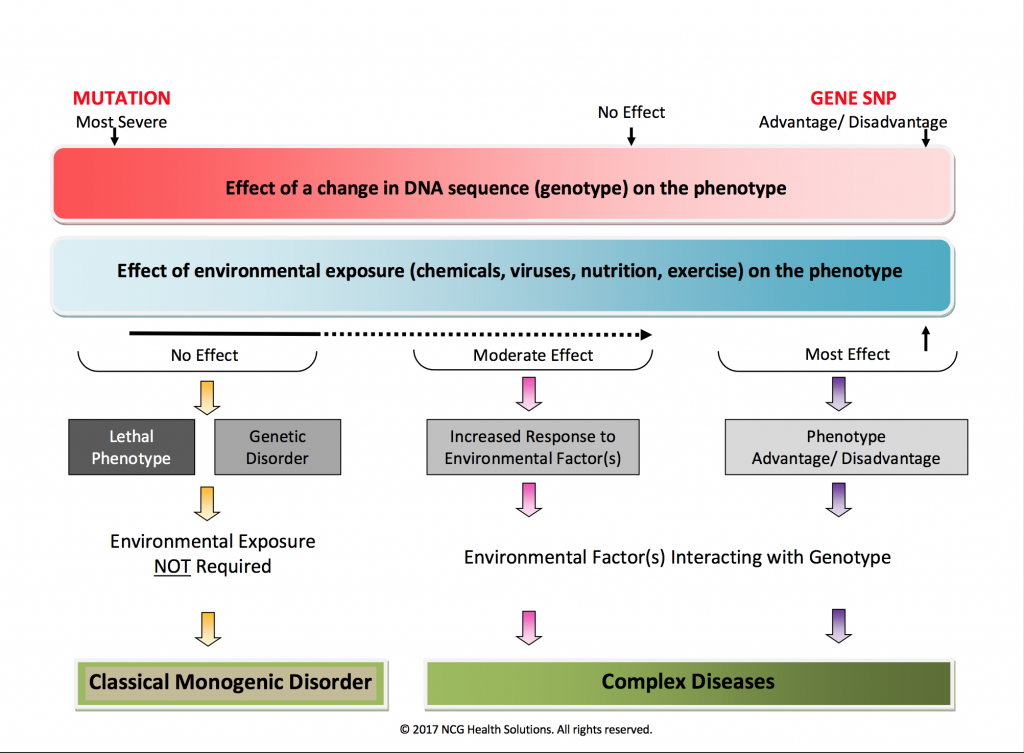
Often, a picture is a worth a thousand words. The FIGURE 1 graphic summarizes the major differences between genetics and genomics. The differences can be gleaned either by going left to right or top to bottom. For example, the red bar at the top depicts the effect of a gene change (genotype) on a person’s phenotype. A mutation, the most severe gene change, is on the left of the graphic, whereas a gene variant or SNP (single nucleotide polymorphism), the most common gene change in the human genome, is represented on the right. A gene SNP can confer a significant advantage to the phenotype or it may be deleterious. Underneath the red bar is a blue bar representing the effect of environmental factors on the phenotype of an individual. The most significant interaction occurs on the far right between one or more gene variants and environmental factors.
FIGURE 1
Reading further down the image, we see that a gene mutation can be lethal or create a classic monogenic disorder. Down Syndrome and cystic fibrosis are two of the more commonly known diseases caused by genetic mutations. As we move across to the right, it gets a little blurry. An example of this is the inherited form of breast cancer caused by BRCA1 and BRCA2 genes. It is classically considered a genetic mutation, and so would belong on the left of the graphic. But, recent studies suggest that the microenvironment associated with the BRCA1 and BRCA2 breast cancer cells may play a role in its development, so we may have to modify this graphic in the future if the early results are confirmed for at least some types of inherited genetic diseases. In general, though, environmental factors do not play a role in the majority of inherited, single-gene disorders.
For nearly every complex disease of the 21st century – including heart disease, arthritis and other autoimmune diseases, cancer, obesity, Alzheimer’s disease, autism, osteoporosis, and more – both genes and environment play a role. A person’s susceptibility to a disease depends on the number of gene SNPs involved and degree of environmental exposure. When multiple gene SNPs confer an increase disease risk, it is referred to as polygenic. This is in contrast to a gene mutation, which is monogenic. We’ll share case histories in future articles highlighting how multiple gene SNPs and estrogen exposure can impact a person’s risk for ER+ breast cancer.
Medical genetics has traditionally focused on health conditions that are due to inherited mutations in single genes (Huntington disease), in whole chromosomes (trisomy 21 or Down syndrome) or are associated with birth defects or developmental disabilities. With a traditional single gene disorder such as Tay-Sachs disease, genetic information is obtained from a patient as well as his or her immediate family members and relatives. The information is then used to diagnosis or predict the risk of an inherited genetic disorder passing from one generation to another via the germ line. For many single gene disorders, there are no or very limited medical interventions available. Although genetic disorders are individually rare, in aggregate they account for about 5% of all human disease.
The remaining 95% are associated with common gene variants that interact with environmental factors to increase or decrease a person’s susceptibility. This is the mission of genomic medicine: uncover how gene variants or SNPs at one or multiple locations interact with environmental factors (diet, drugs, infectious agents, chemicals, and behavioral factors) and influence biochemical pathways, metabolic processes and biological systems, which can lead to the chronic and degenerative diseases associated with aging.
The profound impact of being able to “unzip” a person’s DNA to reduce, reverse or mitigate many chronic diseases was not lost on the early genomic researchers. And to use this tool to construct truly preventive medicine programs has been the promise and dream of genomic medicine. During the past 13 years, genomic testing has expanded beyond the lab bench to the clinic, and continues to grow at an exponential pace.
It is now possible to use results from clinically-based genomic testing companies to evaluate a patient’s disease susceptibility, and develop evidence-based personalized intervention strategies to reduce those risks.
These strategies include patient-specific lifestyle modifications, dietary recommendations, nutritional supplements and/or exercise, which “talk” to genes–that is, up-regulate or down-regulate gene expression. And biomarker testing is used to evaluate whether the intervention is efficacious.
We are often asked is whether this genomics-directed personalized health model is “ready for prime time.” For more than 15 years, clinicians at NCG Health Solutions have been using genomic testing across many different testing platforms with more than 500 proactive individuals, as well as with a cost-conscious corporate wellness program, to direct personalized interventions with great success.
Currently, we use a comprehensive genomic test panel which evaluates 122 genes, with gender-specific reports based on interpretation of more than 1000 gene variants to create personalized programs for the following health conditions:
- prevent ER+ breast cancer or its recurrence in women diagnosed with the condition;
- reverse osteopenia and osteoporosis;
- prevent or treat Type II diabetes, obesity, heart disease and metabolic syndrome;
- optimize sports and exercise performance;
- customize nutrient requirements and resolve nutritional paradoxes;
- prevent, treat and manage depression and anxiety;
- and many more health and wellness issues.
Personalized genomic medicine has reached a tipping point and is revolutionizing healthcare. Be part of the change! Click here to learn more about Genoma International webinars and courses, including the only CME-accredited online courses in genomics and culinary genomics.